Prioritizing Therapeutic Candidates Using Revilico’s scRNA SeqDocumentation Index
Fetch the complete documentation index at: https://docs.revilico.bio/llms.txt
Use this file to discover all available pages before exploring further.
Engine Why Use this Engine? In the documentation below, we will use Revilico’s Single Cell RNA sequencing analysis engine in order to explore and interpret a disease state from a transcriptomic perspective and evaluate whether a particular gene can serve as a potential therapeutic target.
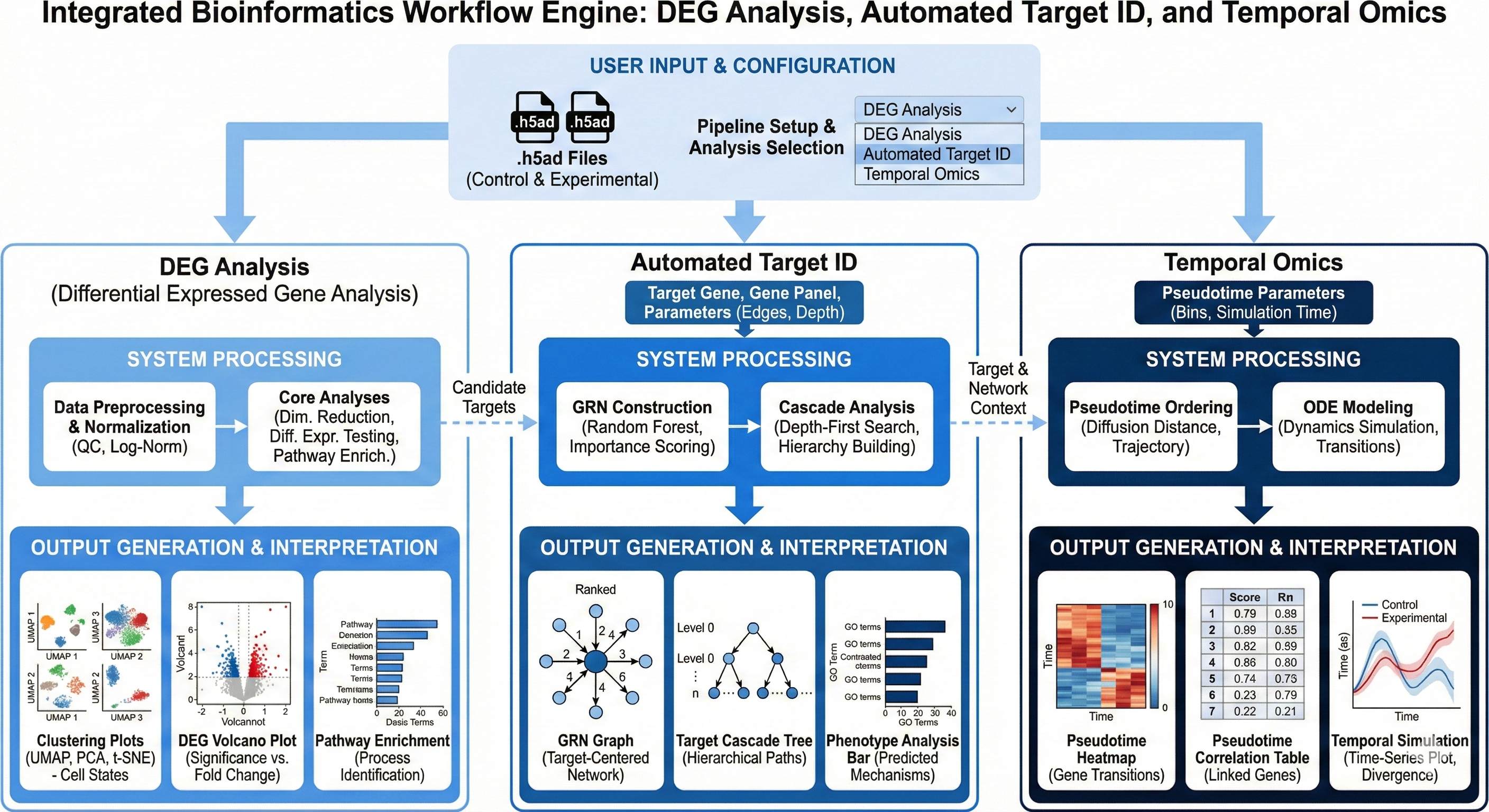
Transcriptomics is the study of an organism’s RNA molecules at a specific time in order to derive insights such as gene expression and gene regulation in many different scenarios ranging from cellular function to comprehension of a disease state. One type of transcriptomic technology in particular is Single Cell RNA sequencing, which is a powerful technique where gene expression of several transcripts can be measured in each individual cell across thousands of cells, allowing users to screen cells of different conditions at a high throughput level. Revilico’s scRNA-seq Analysis Engine integrates 3 different workflows where the user can do the following (1) genome-wide differential expression screening (DEG Analysis) (2) evaluate therapeutic target candidates through gene regulatory network analysis, regulatory correlation mapping, and downstream cascade prediction (Automated Target ID) (3) modeling dynamic gene expression transitions using pseudotime trajectory inference coupled with ODE-based temporal modeling (Temporal Omics Analysis). We will explain these in greater detail below. This is one of our more consolidated engines, so the documentation will be more extensive than other engines. This engine supports hypothesis generation, target validation, and mechanistic understanding of perturbation responses in biological systems. In this documentation, we will use Revilico’s scRNA Seq engine to identify differentially expressed genes, assess the viability of a particular target using Gene Regulatory Network Analyses, and model the temporal dynamics of targeting this particular gene. Below is a diagram of the the workflow for the scRNA seq Engine:

