Documentation Index
Fetch the complete documentation index at: https://docs.revilico.bio/llms.txt
Use this file to discover all available pages before exploring further.
The Problem You are Trying to Solve:
“I want to conduct multiomics analysis with the data I have available to get a better understanding of my desired indication before diving into computational chemistry workflows.”
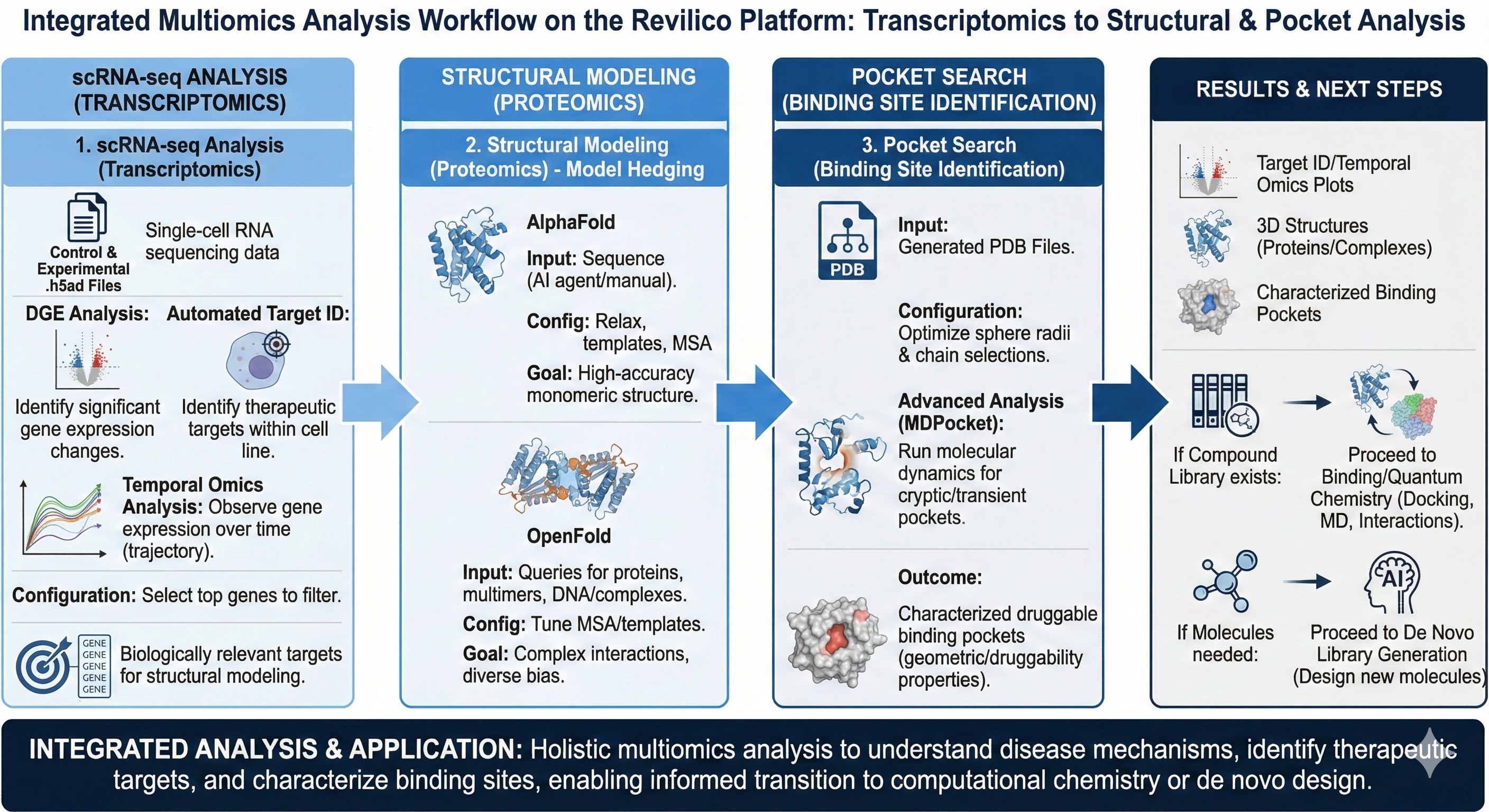 Multiomics analysis is a powerful and holistic approach toward understanding the layers of a biological disease and its mechanisms, including the genomic, transcriptomic, and metabolomic fields. However, conducting multiomics analysis is often challenging, given the various layers and options for interpreting your data. Oftentimes, workflows for multiomics analysis can be scattered or require an individual with bioinformatics/data analytics skills to create custom scripts or pipelines to visualize and analyze results (pipelines that may vary between experiments). Having these models centralized into a unified platform enables users to not only conduct analysis with ease, but to do so through the various layers of this process.
Multiomics analysis is a powerful and holistic approach toward understanding the layers of a biological disease and its mechanisms, including the genomic, transcriptomic, and metabolomic fields. However, conducting multiomics analysis is often challenging, given the various layers and options for interpreting your data. Oftentimes, workflows for multiomics analysis can be scattered or require an individual with bioinformatics/data analytics skills to create custom scripts or pipelines to visualize and analyze results (pipelines that may vary between experiments). Having these models centralized into a unified platform enables users to not only conduct analysis with ease, but to do so through the various layers of this process.
Solution
Here, we present the core features for multiomics analysis currently available on the Revilico platform. Our platform provides extensive workflows and features for transcriptomics and proteomics with additional upcoming features for genomics, metabolomics, and epigenomics in the near future. By using the currently available tools, our hope is that you will be able to have a better understanding of your data and your disease indication before you proceed through the next phases of your drug discovery pipeline.
What Data Do I Need to Provide?
- Control and Experimental h5ad Files (scRNA-seq Analysis)
- CSV with Protein Sequences/Manually Added Protein Sequences (AlphaFold)
- Single Protein, Protein + Ligand, Multimer, or DNA with PTMs (OpenFold)
- PDB Files (Pocket Search)
Workflow
-
scRNA-seq Analysis
To perform transcriptomics analysis by understanding the single-cell RNA sequencing profile of your data, start by uploading both your control and experimental sample data as h5ad files. The platform presents you with three possible analyses that you can select from: 1) DGE Analysis (identify significant changes in gene expression between your sample groups), 2) Automated Target ID (identify ideal targets from your cell line of interest), and 3) Temporal Omics Analysis (observe and analyze the gene expression in your data as a function of time for dynamic and temporal constraints). You can also select the top number of genes you want to extract to filter out the best possible results. Based on the chosen analyses, visualization plots and charts will be available in the results section for this specific tool. This will help you with identification of certain targets of interest using a transcriptomics based approach to move into later stage structure based drug design.
-
AlphaFold
For understanding the structure of your down-selected target/protein, utilize our data extraction AI agents to pull the amino acid sequence you will be utilizing for your protein model, or upload your protein sequence(s) of interest. Here, you have the option to fine-tune the parameters for your protein such as structure relaxation, templates, multiple sequence alignment, and pairing options. Protein structures will be available in the results section for this tool after running.
-
OpenFold
For a greater understanding of the structure of your protein using a different model with different intrinsic biases and performance, and including support for additional complexes, upload your queries based on the data structures accepted. An option is provided to tune multiple sequence alignment or to use templates to help gain more accuracy for your given complexes. 3D structures will be available in the results section for this tool after running.
-
Pocket Search
To identify top pockets of your protein of interest, upload your PDB files into Pocket Search Engine. Here, you have the opportunity to optimize sphere radii and chain selections for your use cases. You can also take this over to the MDPocket tab in this tool to combine your understanding of the available pockets with various molecular dynamic simulations available in our Binding Chemistry suite. Top binding pockets and their characteristics will be available in the results section for this tool after running. These engines help to find static pockets as well as more cryptic or transient pockets that can appear after longer dynamic time scales.
Results
- DGE Analysis, Target ID, Temporal Omics (scRNA-seq Analysis)
- Protein Structures (AlphaFold)
- 3D Structures for Protein, Complexes, or DNA/RNA (OpenFold)
- Top Protein Pockets (Pocket Search)
By running the tools above and performing your analyses, you will be able to get a greater understanding of your disease indication and ideal targets you wish to proceed with. These tools elucidate the underlying biology behind your disease at the cellular pathway level and will enable you to generate structures based on the target or targets identified as potential therapeutic candidates.
Now what?
I have a greater understanding of my disease indication and what target I want to go for!
- If you have molecules or a library of compounds you are interested in using → proceed through the various tools available in our Binding Chemistry or Quantum Chemistry suite to better understand your molecules’ binding, docking, molecular dynamics, or quantum-level interactions.
- If you do not have molecules or a library of compounds → proceed to the De Novo Library Generation available in our Binding Chemistry suite to develop molecules for your newly selected target. You can also reference our pre-determined libraries that are on hand in liquid and power form.
Why Revilico?
This Revilico workflow enables users to query the availability of experimental data before proceeding to AI structure prediction. All 3D structural hypotheses include transparent quality scoring, and can be seamlessly integrated with downstream discovery engines and workflows in a multi-modal way to help hedge all of your results against one another.

