Documentation Index
Fetch the complete documentation index at: https://docs.revilico.bio/llms.txt
Use this file to discover all available pages before exploring further.
Why use this engine?
In the documentation below, we will use Revilico’s Pocket Search Engine to identify potential pockets to target with drug binding. Using Pocket search we will identify pockets on the static version of the protein based simply on the geometry of the protein and use MDPocket to simulate which pockets remain stable while the protein is in motion. This portion of computational chemistry is essential to understanding how key biological functions can be enhanced or blocked using different therapeutic strategies. This algorithm mainly searches for druggability and chemistry, and when supplemented with prior biological knowledge and literature review, it can be a powerful tool to regulate biologies.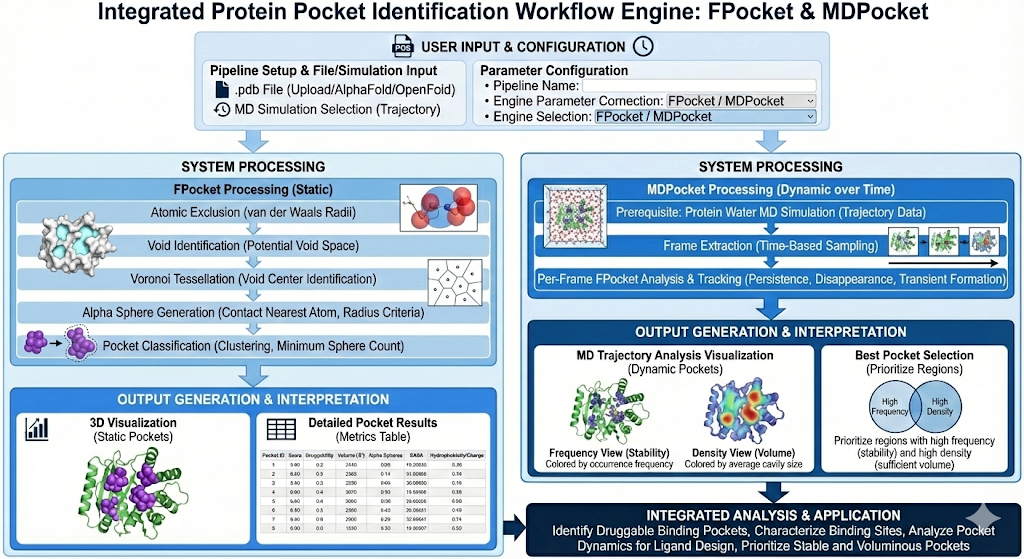
Background
Pocket identification is a critical aspect of drug discovery. It allows researchers to understand protein function and enable rational drug design by pinpointing specific areas along the protein where small molecules, ligands, or other proteins can bind. In Revilico’s Pocket Search Engine, we introduce two tools, one for understanding the protein from a static perspective and one for understanding the protein from a dynamic perspective. Our pocket search engine is a purely geometry based algorithm that detects binding pockets by analyzing protein shape and cavity architecture. We will now dive into the theory behind the algorithm on which the system was built on. We first start with our protein. On each atom of this protein, we assign an atomic exclusion zone defined by its van der Waals (dW) radius. The circumference of this circle defines the boundaries of this atom, and for each element there will be an assigned radius in Angstroms where no ligand atom can exist. We then define all other space not in this atomic exclusion zone as potential void space in which a ligand atom can exist. The next step is understanding within this void where we can identify these void centers. This void center identification is based on Voronoi tessellation which is a mathematical framework defined by the following equation: For a set of protein atoms , the Voronoi cell of atom is Where p is any point in 3D space. Essentially what this means is that a Voronoi cell contains all points closer to Ai than to any other atom. Where we identify a void center is that this is a point that is equidistant from typically 4 or sometimes more voronoi centers. We then generate a sphere with the center located at this void center, and expand until it touches the surface of the nearest atom. If the sphere generated is greater than our min alpha sphere radius and less than our max alpha sphere radius. It remains a valid alpha sphere. The min and max of the sphere sizes are pre-determined. Once multiple valid spheres are generated, if we meet our min spheres per pocket parameter within our clustering distance, it is then classified as a pocket. Another way to look at this equation and system is that we are able to ‘roll a sphere’ across the entire protein and detect the chemistries along that sphere’s route to be able to extract critical parameters that we can use in downstream assessments. Identifying Pockets with Revilico’s Pocket Search EnginePocket search is an engine where the primary goal is to answer the question: Where are the physical cavities large enough to accommodate drug-like molecules. This engine is essential for identifying potential binding sites for drug design, rank pockets by druggability before expensive docking calculations, mapping surface topology for understanding protein function, selecting docking regions based on geometric feasibility, and generating hypotheses for mutagenesis or fragment screening. Usually, you can use literature based on experimental methods to determine regions on the amino acid chains that contribute to biological activity to overlay with these pocket search algorithm regions to confirm or identify pockets. Identifying Pockets with Revilico’s MDPocket Engine
MD Pocket operates the same way as Pocket search, the difference being that it calculates the Pocket search at every frame of a MD simulation, capturing the pockets while the protein dynamically changes, compared to Pocket search which just captures the pockets at one screenshot of the protein. MDPocket reveals which pockets persist, disappear, or transiently form over time.
Interactive Pocket Viewer
Explore FPocket pocket detection results in an interactive 3D viewer. The protein structure is shown at 50% transparency with detected pockets highlighted as spacefill overlays.- All Pockets View — See all detected pockets overlaid in red on the protein structure.
- Individual Pocket — Select a specific pocket to highlight it in orange with detailed statistics.
- Pocket Statistics — View druggability score, volume, SASA, hydrophobicity, and more for each pocket.

