Documentation Index
Fetch the complete documentation index at: https://docs.revilico.bio/llms.txt
Use this file to discover all available pages before exploring further.
Why Use this product?
The BoltzGen Cofolding Engine is a diffusion based tool that generates thousands of candidate binders with validated binding poses enabling rapid discovery of high affinity binders for drug development, diagnostics, or protein engineering applications without requiring experimental screening. This tool is best used when you need to design novel protein or peptide binders against therapeutic targets through AI powered de novo generation that simultaneously optimizes binding affinity, structural stability, and sequence diversity.
Background
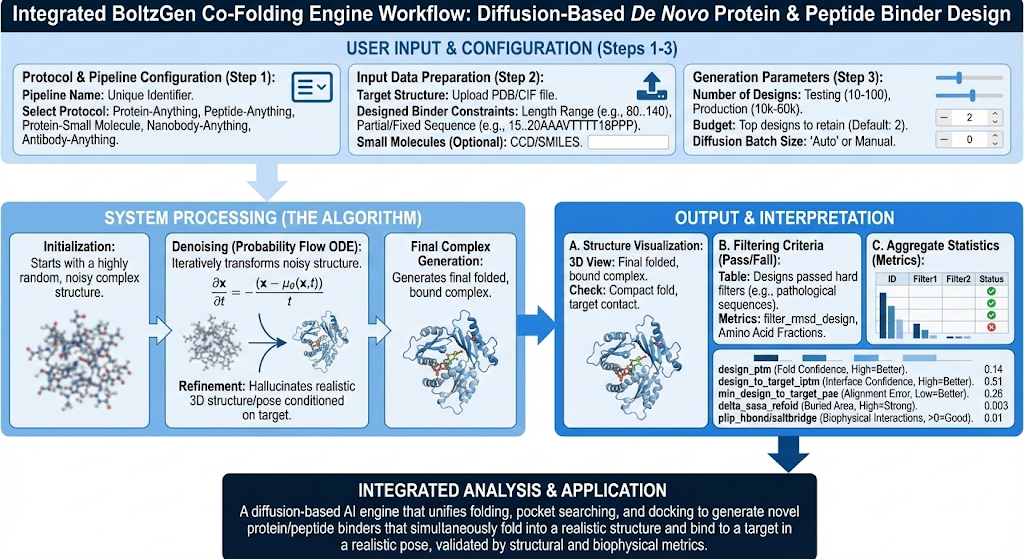
Typical workflows for discovering whether a molecule will bind to a protein pocket are a multistep process that is both time consuming and computationally expensive. And even with these workflows it is not guaranteed that the pocket we select is the correct pocket. BoltzGen Co-Folding is a tool that can be deployed to design a new binder that will both fold into a realistic 3D structure and bind to a target in a realistic 3D pose, in one unified generative process. This process can bypass the usual process of protein folding, then pocket search, then docking for traditional small molecule therapeutic developments.
BoltzGen Co-Folding works by uploading a target structure, typically a fixed protein structure as a PDB file, then design constraints for the binder in which we want to create (e.g. 80..140 is a length constraint for the number of residues). BoltzGen will then take these two structures and build one joint complex where we have the fixed target, and the designed binder (i.e. small molecule, peptide, etc.). BoltzGen is usually utilized for designing biologics modalities for therapeutics against specific targets.
BoltzGen will then run a generative model that is an all-atom diffusion style generative model, where designed entities are represented at the atomic level, grouped into tokens (i.e. residues/nucleotides), while designed parts are represented with naked residue/atom types in a fixed-size representation. The model will take in an input of known atoms/coordinates for the target, tokens for all residues (target + binder), with masks indicating which residue/atom types are designable, and conditioning signals based on the design constraints we have provided in the input. It will output a generated all-atom 3D structure for the whole complex, where a denoiser prediction will be used to iteratively refine the sample. In order words, the model is sampling a full complex from a learned distribution to see what real complexes look like, conditioned on your target and constraints. It can be denoted by the following equation:
P(sequence,structure,pose∣fixed anchor,constraints)
The model starts from a highly random complex and iteratively transforms it into a more statistically realistic protein-target complex, according to patterns learned from real structures. This concept of a diffusion style generative model takes on the idea of taking a protein complex adding noise until it looks like random junk, then training a neural network to undo this noise towards an optimized 3D conformation and engagement. The goal is to create a model that can reliably clean up noise resulting in a realistic structure. This can be denoted by the following equation:
Dθ(xt,t,conditioning)
Where xt is the noisy version of a structure at step t, t is how noisy it is, and conditioning is the fixed protein, ligand, constraints. Our output will be a less noisy structure.
At the generation step BoltzGen will use a probability flow ODE to generate the new complex. It can be denoted by the following equation:
∂t∂x=−tx−μθ(x,t)
Where X is the current molecular configuration at a given state, t is a continuous parameter controlling how noise the structure is with large t indicating very noisy random structure and small t indicating low noise and close to a final realistic structure, μθ(x,t) is the model’s denoised prediction which represents the model’s best guess of what this structure would look like if it were clean and realistic, and dx/dt indicating the direction of change of how to update the structure as noise is reduced.
To summarize, this model will deploy design steps to iteratively transform a random joint structure into a more statistically realistic bound complex, where the design constraints and the presence of the fixed target shape what ‘realistic’ means at every step
Interactive Results Viewer
Explore BoltzGen Co-Folding results interactively. View generated protein designs with filtering criteria, aggregate statistics, ranked structures, and metrics visualization.


