Documentation Index
Fetch the complete documentation index at: https://docs.revilico.bio/llms.txt
Use this file to discover all available pages before exploring further.
The Problem You Are Trying to Solve
“I have a set of lead compounds, and I want to understand both their direct on-target effects and their indirect off-target effects, including how they engage broader biological pathways and cellular programs.”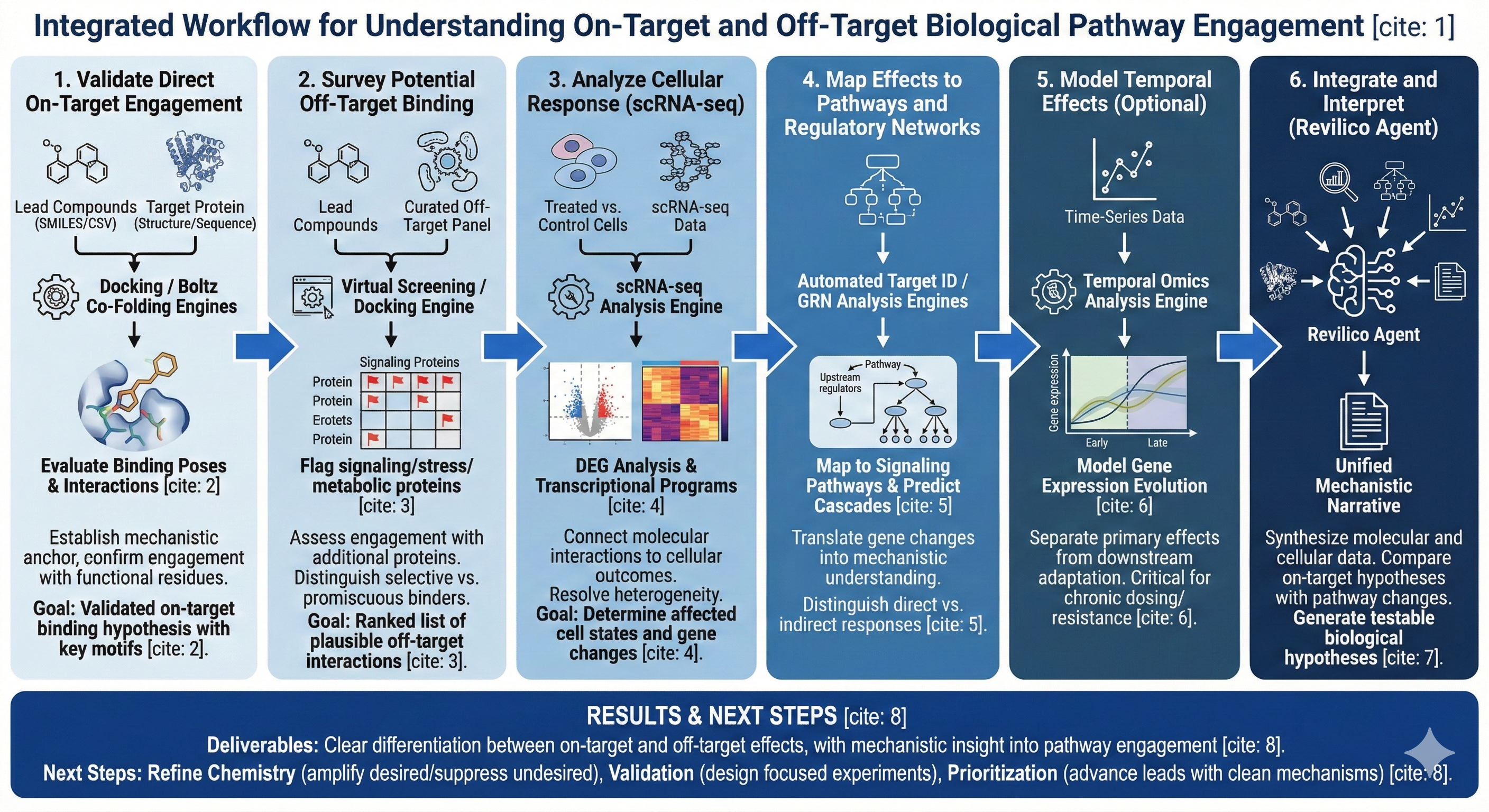
- Whether the compound robustly engages the intended target in a biologically meaningful way
- Which secondary proteins or pathways may be perturbed downstream
- Whether observed phenotypes arise from on-target mechanism, off-target liabilities, or network-level effects
- How molecular binding events translate into cellular state changes
Solution
This workflow integrates binding chemistry engines with transcriptomic pathway analysis to connect molecular engagement with biological consequence. The primary analysis chain is: On-Target Binding Validation → Off-Target Binding Survey → Transcriptomic Response Analysis → Pathway & Network Interpretation. This allows users to:- Confirm intended target engagement
- Identify plausible off-target interactions
- Observe how cells respond to compound treatment
- Resolve whether effects are direct, indirect, or compensatory
What Data Do I Need to Provide?
Required- Lead compound structures (SMILES or CSV)
- Primary target protein structure or sequence
- Cellular transcriptomic data (e.g., scRNA-seq) from treated vs control conditions
- Time-resolved transcriptomic data (multiple doses or timepoints)
- Known pathway annotations for the disease context
- Off-target protein panels
- Comparative compounds (tool compounds, inactive controls)
Workflow
- Validate Direct On-Target Engagement
Using Docking and (if applicable) Boltz or BoltzGen Co-Folding, users:
- Evaluate binding poses and interaction patterns
- Confirm engagement of known functional residues
- Compare multiple leads for consistency of on-target binding
- Survey Potential Off-Target Binding
- Identify secondary binding candidates
- Flag proteins involved in signaling, metabolism, or stress responses
- Distinguish selective compounds from promiscuous binders
- Analyze Cellular Response with scRNA-seq
- Perform differential gene expression (DEG) analysis between treated and control cells
- Identify transcriptional programs altered by compound exposure
- Resolve heterogeneity across cell populations
- What changes when the compound is applied?
- Which cell states are most affected?
- Map Effects to Pathways and Gene Regulatory Networks
- Identify upstream regulators driving observed transcriptional changes
- Map altered genes to known signaling pathways
- Predict cascade effects downstream of target engagement
- Direct on-target pathway modulation
- Indirect off-target or compensatory responses
- Model Temporal and Dose-Dependent Effects (Optional)
- Model how gene expression evolves after compound exposure
- Identify early vs late response programs
- Separate primary effects from downstream adaptation
- Chronic dosing scenarios
- Pathway rewiring and resistance studies
- Integrate and Interpret with Revilico Agent
- Ask mechanistic “why” questions across datasets
- Compare on-target binding hypotheses with observed pathway changes
- Generate testable biological hypotheses for follow-up experiments
Results
- Clear differentiation between on-target and off-target effects
- Mechanistic insight into pathway-level engagement
- Reduced ambiguity in phenotypic interpretation
- Stronger confidence in lead progression or redesign decisions
Now What? I understand how my leads engage biology, but what’s next?
Common next steps include:- Refining chemistry to amplify desired pathways and suppress undesired ones
- Designing focused validation experiments
- Prioritizing leads with clean, interpretable mechanisms
- Integrating findings into toxicity or efficacy optimization workflows
Why Revilico?
Revilico uniquely connects:- Molecular binding engines for direct interaction analysis
- Single-cell transcriptomics for biological response resolution
- Network and temporal modeling for pathway-level insight
- AI-assisted reasoning to unify complex datasets

