Documentation Index
Fetch the complete documentation index at: https://docs.revilico.bio/llms.txt
Use this file to discover all available pages before exploring further.
The Problem You are Trying to Solve:
“I have a drug with demonstrated efficacy but unknown mechanism of action, and I want to identify its primary molecular target(s) to enable structure-based optimization and rational drug design”
Traditional target identification for phenotypic hits required labor-intensive biochemical approaches like affinity chromatography pull-down assays, proteomics screens, and genetic validation studies, each taking months and often yielding ambiguous results with multiple potential targets. Without knowing the binding site or target protein, rational optimization can be impossible, forcing researchers into costly trial and error medicinal chemistry campaigns that modify the drug blindly while monitoring phenotypic readouts. The disconnect between observable efficacy and molecular mechanism meant many promising drugs were abandoned or progressed to clinical trials without understanding their liabilities, leading to unexpected toxicities or failure to translate across disease contexts. Moreover, during the acquisition process of therapeutic candidates, knowing the target of interest is exceptionally important as it allows for a fall back to structure based drug design should anything go wrong during clinical trials or beyond.
Solution
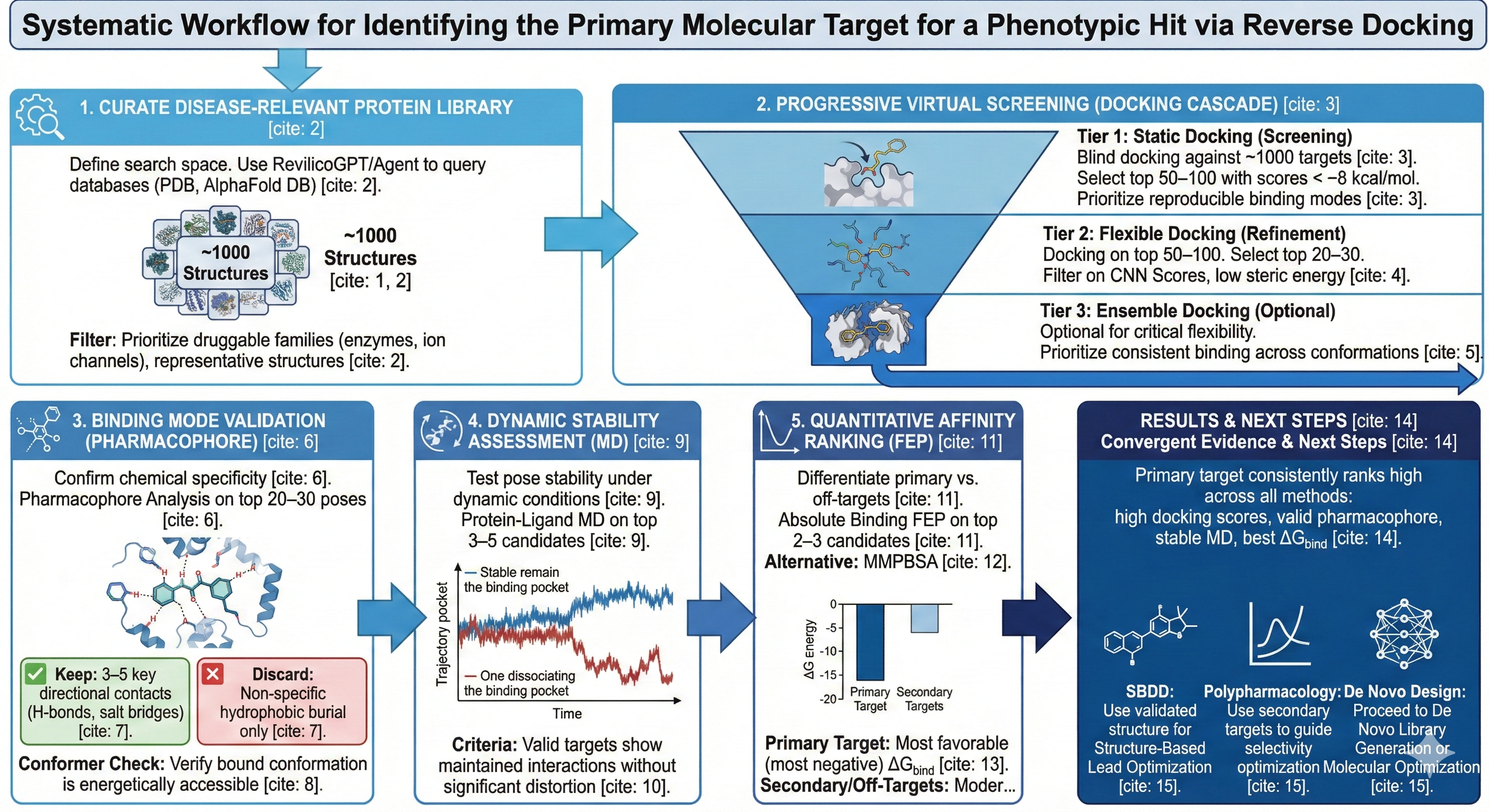
This workflow enables users to a) systematically identify primary molecular targets for phenotypic hits through reverse docking against disease-relevant protein libraries and b) validate target engagement through progressive refinement using Revico’s multi-tiered virtual screening and molecular dynamics engines. We leverage a cascade of increasingly rigorous computation methods, i.e. static docking for rapid screening, flexible docking for improved accuracy, ensemble docking for conformation sampling, protein-ligand MD for dynamic stability assessment, and free energy perturbation for quantitative affinity ranking at the highest level of accuracy. Proper target engagement validation is achieved through pharmacophore analysis that confirms reasonable binding modes and binding free energy calculations that distinguish true targets from docking artifacts.
What Data Do I Need to Provide?
- Known ligand or binding partner (required for docking, MD, FEP, and Pharmacophore analysis)
- Protein Library (Optional, we have libraries on hand, or we can generate this using RevilicoGPT/Revilico Agent as well for targeted protein sets)
Workflow
- Curate Disease-Relevant Protein Library
Assemble a comprehensive library of protein structures associated with your therapeutic context to define the search space for reverse docking. Use RevilicoGPT/Revilico Agent to query structural databases (PDb, AlphaFold Database) and filter for disease relevant targets
Sample Query: I have a drug that shows efficacy in [disease/phenotype]. Please compile a library of ~1000 protein structures associated with this disease, prioritizing druggable targets like kinases, GPCRs, enzymes and ion channels. Return PDB files, amino acid sequences, or AlphaFold structures with binding site annotations.
Revilico Agent will output a curated protein structure library with metadata including protein family, known functions, disease associations, and druggability scores. Users can refine this library by filtering for specific protein families or expanding to the full druggable proteome for unbiased screening. The library should include experimentally resolved structures when available and high-confidence AlphaFold predictions to maximize coverage. For proteins with multiple conformations available, include representative structures capturing different functional states to account for conformational selectivity
- Progressive Virtual Screening via Multi-Tiered Docking
Screen your drug molecules against the protein library using a cascade of increasingly rigorous docking methods to efficiently identify and refine target candidates. Start with rapid screening to eliminate non-binders, then apply more accurate methods to top candidates.
Run Static Docking against all ~1000 proteins in your library. This high-throughput screen rapidly eliminates non-binders based on binding affinity predictions. This can be done through ‘blind docking methods’ where the entire protein surface is exposed to the algorithm to calculate binding energies, placing the compound in what it believes to be the best suited position. Downselect to top 50-100 proteins with binding affinities < -8 kcal/mol, prioritizing targets with multiple low-energy poses in consistent binding sites, which indicates reproducible binding modes rather than spurious predictions
Run Flexible Docking on the top 50-100 candidates from static docking. This will account for side-chain flexibility in the binding site, providing more realistic binding geometries and hybrid physics ML scoring. Downselect to the top 20-30 proteins based on CNN Pose Score, CNN Affinity, and low intramolecular (sterics) energy.
For targets where protein flexibility is critical, you can optionally run Ensemble Docking using MD-derived conformational ensembles. This captures how your drug binds across the protein’s conformational landscape. Prioritize targets showing consistent binding across multiple protein conformations over those with conformation-dependent binding.
- Validate Binding Modes with Pharmacophore Analysis
For the top 20-30 candidates from flexible docking, analyze the binding interactions to confirm chemically reasonable binding modes before committing to expensive MD simulations. Run Pharmacophore Analysis on the top-ranked docked poses. Prioritize targets where the drug forms multiple complementary interactions (typically 3-5 key contacts). Filter out targets where binding relies solely on hydrophobic burial with no directional interaction. Use Conformer Search on your drug to validate that the bound conformation is energetically accessible to ensure the drug doesn’t pay a large conformational penalty for binding.
- Dynamic Stability Assessment via Molecular Dynamics
Run Protein-Ligand MD simulations for your top 3-5 target candidates. MD reveals whether static docking purposes remain stable or dissociate, whether key interactions persist, and whether the protein binding site accommodates the drug without creating a strain. Strong candidates show stable complex formation, maintained binding interactions, and no signs of ligand egress or binding site distortion.
- Quantitative Affinity Ranking via Free Energy Calculations
For your top 2-3 candidates, calculate absolute binding free energies to quantitatively rank targets and identify the primary binding partner versus secondary off-targets. The target with the most favorable (most negative) ΔG binding is your primary molecular target, while targets with weaker binding may represent secondary pharmacology or off targets.. Consider running MMPBSA analysis from MD trajectories as a faster less rigorous alternative if FEP is computationally expensive and costs too many credits for your campaign.
Results
- Ranked candidate target list with binding affinities and pose clusters
- Pharmacophore interaction maps (e.g. H-bonds, salt bridges, key contacts)
- MD stability metrics (RMSD, RMSF, interaction persistence)
- Absolute binding free energies (ΔG bind) per target
- Primary target identification with validated binding mode
Look for convergent evidence across methods, your primary target should rank consistently high in docking scores, show 3-5 chemically reasonable interactions in pharmacophore analysis, maintain stable binding in MD simulations, and demonstrate the most favorable ΔG bind in FEP calculations. Use the validated primary target structure and binding mode to proceed with structure based lead optimization, while documenting secondary targets with moderate affinity for selectivity optimization or polypharmacology consideration. You should also keep in mind that your compound could be engaging with and interacting with several targets all at once, so you need to overlay your results with biologically feasible pathways that you believe are being affected during phenotypic screening. This helps to overlay and connect the dots between binding engagements and biological outcomes.
Now what? You may have your target of interest now (after you have validated it experimentally) and may want to run optimization to get tighter biochemical engagements
- After analyzing and down-selecting your targets of interest, you can then move into generative chemistry campaigns using De Novo Library Generation, Molecular Optimization, or Custom Model training to get new libraries optimized for engagement to your target.
- After getting results from generative chemistry, you can re-score the new library using different tools to get and expand your lead series to be synthesized and tested.
Why Revilico?
This workflow addresses the challenges of identifying the primary molecular target for a drug with demonstrated efficacy but an unknown mechanism of action, which is critical for rational structure based optimizations. This process systematically identifies and validates protein targets of interest before moving forward with costly experimental validations like siRNA knockdowns or CRISPR screens.