Documentation Index
Fetch the complete documentation index at: https://docs.revilico.bio/llms.txt
Use this file to discover all available pages before exploring further.
The Problem You are Trying to Solve:
“I have a molecular target without obvious or stable binding sites, and I want to identify druggable pockets suitable for structure based drug design”
Traditionally identifying binding sites on “undruggable” targets required expensive experimental methods like fragment screening via X-ray crystallography, NMR spectroscopy to detect conformational dynamics, or hydrogen-deuterium exchange mass spectrometry to identify flexible regions, all requiring months of work and significant material costs with no guarantee of finding a druggable site.
Solution
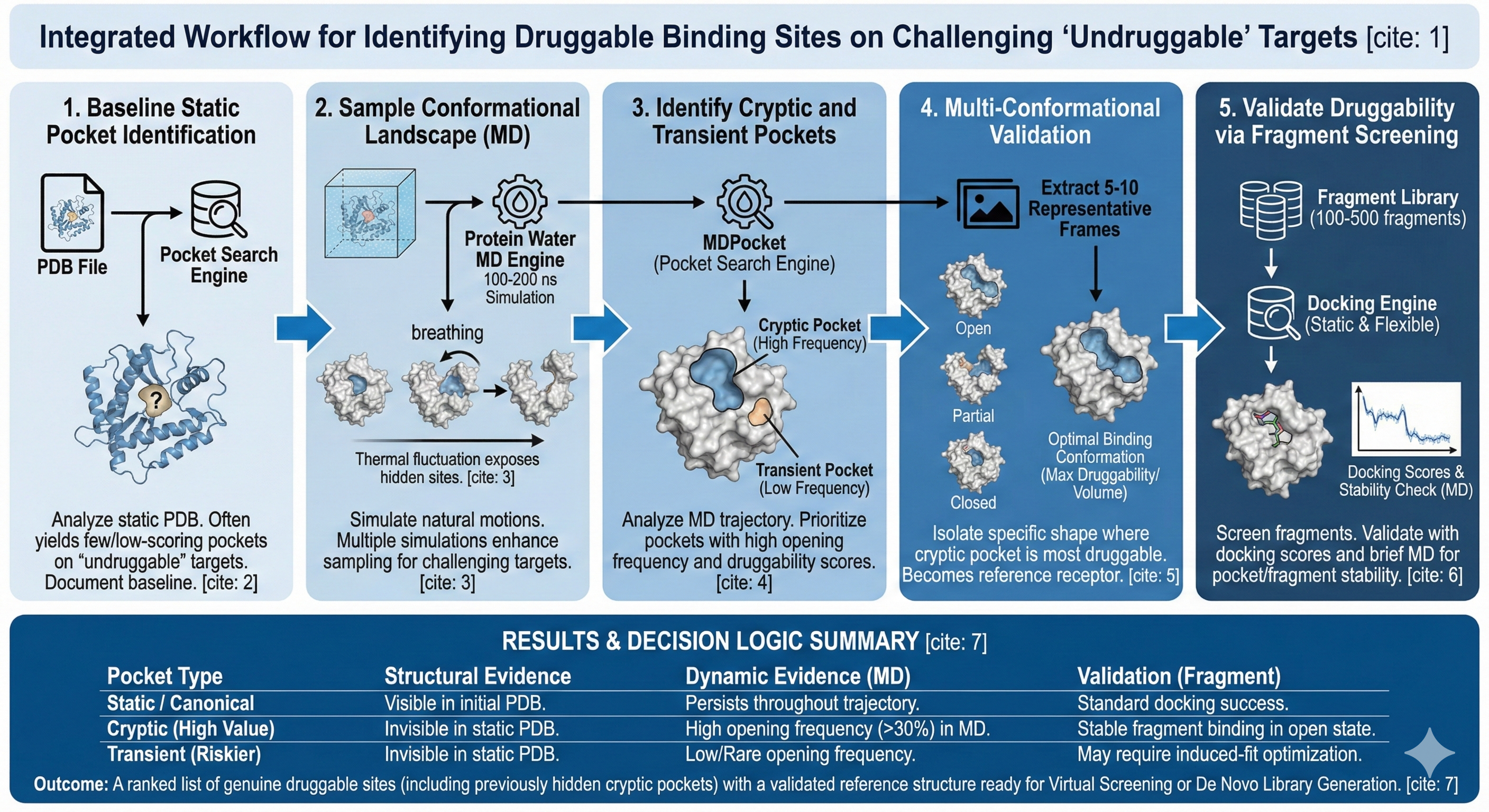
This workflow enables users to a) systematically identify all potential binding sites on challenging targets through static pocket analysis and b) discover cryptic, transient, and allosteric pockets that only appear during protein motion using molecular dynamics-based pocket detection across Revilio’s structural analysis engines. Proper druggability assessment is achieved by analyzing pocket properties across multiple protein states and ranking sites by persistence, accessibility, and druggability scores, helping eliminate status analysis blind spots and increasing confidence that identified pockets represent genuine druggable sites even on targets traditionally considered undruggable.
What Data Do I Need to Provide?
- PDB File of your protein structure (will be used for Pocket Search, Docking, and MD simulations)
Workflow
- Baseline Static Pocket Identification
Run Pocket Search Engine on your protein structure. If you have multiple conformational states, run the analysis on each independently after extracting trajectory .pdb files. This engine will identify cavities and rank them by druggability score, volume, and physicochemical properties. For traditionally undruggable targets, you may find a few pockets or only low scoring ones at this stage. Document all identified pockets with their location as these provide context for interpreting cryptic pockets discovered later.
- Sample Conformational Landscape via Molecular Dynamics
Run Protein-Water MD simulations with explicit solvent for 100-200 ns. The simulation allows the protein to explore thermally accessible conformational states under physiological conditions. During the trajectory, pockets will open, close, expand, contract, and new cavities will transparently appear as the protein breathes and fluctuates. The MD trajectory becomes the input for temporal pocket analysis in Step 3. For particularly challenging targets, consider running multiple independent MD simulations starting from different initial conformations to enhance conformational sampling and increase probability of discovering rare cryptic pocket opening events. You can also explore new configurations of the solvent as well as longer time scales to allow the proteins to reach equilibrium.
- Identify Cryptic and Transient Pockets
Run MDPocket on the MD trajectory from Step 2. MD pocket performs pocket detection at each trajectory frame and tracks pocket properties over time identifying cryptic pockets, transient pockets, persistent pockets, and pocket frequency. MDPocket will output ranked cryptic pockets with temporal profiles showing when each pocket opens, how long it remains accessible, and its properties over time. Cryptic pockets with high opening frequency and favorable druggability represent prime targets for allosteric or induced-fit drug design strategies that capitalize on protein flexibility.
- Multi Conformational Validation
Extract representative protein conformations from the MD trajectory and validate pocket characteristics in specific functional states to confirm MDPocket findings and understand pocket architecture. From the MD trajectory, extract 5-10 representative conformations showing different pocket states: cryptic pockets in fully open, partially open, and closed states. For more extensive analysis, you can cluster trajectory frames by backbone RMSD/RMSF or pocket volume to identify structurally distinct conformations. Run Pocket Search Engine on each extracted conformation independently. Compare pocket properties across conformations to understand how pocket characteristics change with protein state. Identify the “optimal binding conformation” where the cryptic pocket exhibits maximum druggability. This becomes the reference structure for subsequent structure based drug design and optimization studies.
- Validate Druggability with Fragment Screening
Run Static Docking and Flexible Docking with a diverse fragment library (around 100-500 small fragments), into the top-ranked cryptic pockets identified in steps 3-4. Use the optimal binding conformation extracted in step 4 as the receptor structure, or perform ensemble docking across multiple conformations if the pocket shows conformational flexibility. For the top fragment hits in cryptic pockets, run brief Protein-Ligand MD simulations to confirm the pocket remains stable when occupied and the fragment doesn’t induce pocket collapse or protein unfolding. Successful fragment binding validates that the cryptic pocket is a genuine druggable site suitable for hit-to-lead optimization. We will soon be coming out with an engine that allows for pocket search using fragment ‘soup’ that will allow us to run longer term molecular dynamic trajectories in the presence of fragment perturbations to elucidate unique pockets of interest.
Results
- Comprehensive pocket inventory with druggability scores and volumes
- Temporal pocket profiles showing opening frequency and persistence across MD simulations
- Representative protein conformations with pocket open/closed states
- Fragment screening validation results
- Ranked druggability site recommendations with strategic classifications
Prioritize cryptic pockets with convergent evidence: high opening frequency, favorable druggability scores, successful fragment binding validation, and maintained stability in ligand-bound MD simulations. These represent genuine druggable sites on previously undruggable targets. If dynamic analysis reveals persistent cryptic pockets that were invisible in static structures and validate with fragment screening, you’ve successfully identified tractable binding sites for structure-based drug design
Now what? After identifying your pockets of interest, you can then move into identifying hit molecules for your campaign.
- Utilize the Virtual Screening engine or De Novo Library Generation to begin your campaign and identify initial hits. Utilizing the knowledge gained from pocket identification, you can target your docking calculations on the box of choice and across conformational states to ensure you’re representing the biological system properly.
Why Revilico?
This workflow addresses the challenge of identifying druggable sites on targets without obvious or stable binding pockets by combining static analysis with dynamic molecular dynamics (MD) simulations. It systematically discovers cryptic and transient binding sites through MD and MDPocket, then validates the most persistent and druggable pockets using fragment screening to establish genuine targets for downstream structure-based drug design.